Drag the Terms to Their Corresponding Class in Order to Review Various Aspects of Gene Regulation
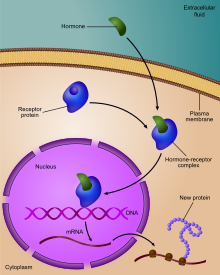
Regulation of gene expression past a hormone receptor
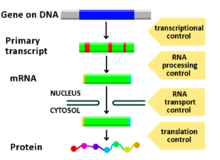
Diagram showing at which stages in the Dna-mRNA-protein pathway expression can be controlled
Regulation of gene expression, or gene regulation,[ane] includes a wide range of mechanisms that are used by cells to increase or decrease the product of specific factor products (poly peptide or RNA). Sophisticated programs of gene expression are widely observed in biology, for example to trigger developmental pathways, respond to environmental stimuli, or conform to new food sources. Well-nigh whatsoever step of factor expression can be modulated, from transcriptional initiation, to RNA processing, and to the post-translational modification of a protein. Oft, one gene regulator controls another, then on, in a factor regulatory network.
Gene regulation is essential for viruses, prokaryotes and eukaryotes every bit it increases the versatility and adjustability of an organism by allowing the prison cell to limited protein when needed. Although as early every bit 1951, Barbara McClintock showed interaction betwixt two genetic loci, Activator (Ac) and Dissociator (Ds), in the color germination of maize seeds, the kickoff discovery of a gene regulation system is widely considered to be the identification in 1961 of the lac operon, discovered by François Jacob and Jacques Monod, in which some enzymes involved in lactose metabolism are expressed by E. coli just in the presence of lactose and absence of glucose.
In multicellular organisms, gene regulation drives cellular differentiation and morphogenesis in the embryo, leading to the creation of dissimilar jail cell types that possess different gene expression profiles from the aforementioned genome sequence. Although this does not explicate how gene regulation originated, evolutionary biologists include it equally a partial caption of how evolution works at a molecular level, and information technology is key to the scientific discipline of evolutionary developmental biology ("evo-devo").
Regulated stages of factor expression [edit]
Any step of factor expression may be modulated, from signaling to transcription to post-translational modification of a protein. The following is a list of stages where gene expression is regulated, the most extensively utilized point is Transcription Initiation:
- Bespeak transduction
- Chromatin, chromatin remodeling, chromatin domains
- Transcription
- Post-transcriptional modification
- RNA send
- Translation
- mRNA degradation
Modification of DNA [edit]
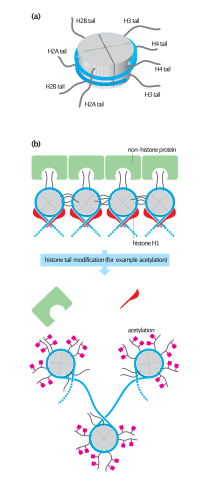
Histone tails and their function in chromatin formation
In eukaryotes, the accessibility of big regions of Deoxyribonucleic acid can depend on its chromatin structure, which tin exist altered as a result of histone modifications directed past DNA methylation, ncRNA, or Dna-bounden poly peptide. Hence these modifications may up or downwardly regulate the expression of a gene. Some of these modifications that regulate factor expression are inheritable and are referred to as epigenetic regulation.
Structural [edit]
Transcription of DNA is dictated past its construction. In general, the density of its packing is indicative of the frequency of transcription. Octameric protein complexes called histones together with a segment of DNA wound around the viii histone proteins (together referred to as a nucleosome) are responsible for the amount of supercoiling of DNA, and these complexes tin can exist temporarily modified past processes such every bit phosphorylation or more than permanently modified by processes such as methylation. Such modifications are considered to be responsible for more than or less permanent changes in gene expression levels.[2]
Chemical [edit]
Methylation of Dna is a common method of gene silencing. DNA is typically methylated by methyltransferase enzymes on cytosine nucleotides in a CpG dinucleotide sequence (as well called "CpG islands" when densely amassed). Assay of the blueprint of methylation in a given region of DNA (which tin can exist a promoter) tin be achieved through a method called bisulfite mapping. Methylated cytosine residues are unchanged by the handling, whereas unmethylated ones are changed to uracil. The differences are analyzed past DNA sequencing or by methods developed to quantify SNPs, such as Pyrosequencing (Biotage) or MassArray (Sequenom), measuring the relative amounts of C/T at the CG dinucleotide. Aberrant methylation patterns are idea to be involved in oncogenesis.[3]
Histone acetylation is also an important process in transcription. Histone acetyltransferase enzymes (HATs) such equally CREB-bounden poly peptide also dissociate the Dna from the histone complex, allowing transcription to proceed. Often, Deoxyribonucleic acid methylation and histone deacetylation piece of work together in gene silencing. The combination of the two seems to be a indicate for DNA to exist packed more densely, lowering cistron expression.[ citation needed ]
Regulation of transcription [edit]
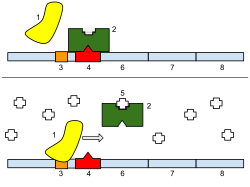
1: RNA Polymerase, 2: Repressor, iii: Promoter, 4: Operator, 5: Lactose, 6: lacZ, 7: lacY, viii: lacA. Top: The factor is essentially turned off. In that location is no lactose to inhibit the repressor, so the repressor binds to the operator, which obstructs the RNA polymerase from binding to the promoter and making lactase. Bottom: The gene is turned on. Lactose is inhibiting the repressor, allowing the RNA polymerase to bind with the promoter, and express the genes, which synthesize lactase. Eventually, the lactase will digest all of the lactose, until in that location is none to bind to the repressor. The repressor will and then bind to the operator, stopping the manufacture of lactase.
Regulation of transcription thus controls when transcription occurs and how much RNA is created. Transcription of a cistron by RNA polymerase can exist regulated past several mechanisms. Specificity factors alter the specificity of RNA polymerase for a given promoter or fix of promoters, making it more or less probable to demark to them (i.east., sigma factors used in prokaryotic transcription). Repressors bind to the Operator, coding sequences on the DNA strand that are close to or overlapping the promoter region, impeding RNA polymerase'due south progress forth the strand, thus impeding the expression of the cistron. The image to the right demonstrates regulation by a repressor in the lac operon. General transcription factors position RNA polymerase at the start of a protein-coding sequence and then release the polymerase to transcribe the mRNA. Activators enhance the interaction between RNA polymerase and a detail promoter, encouraging the expression of the gene. Activators exercise this past increasing the attraction of RNA polymerase for the promoter, through interactions with subunits of the RNA polymerase or indirectly past changing the construction of the Deoxyribonucleic acid. Enhancers are sites on the Deoxyribonucleic acid helix that are jump by activators in club to loop the DNA bringing a specific promoter to the initiation circuitous. Enhancers are much more common in eukaryotes than prokaryotes, where simply a few examples exist (to date).[iv] Silencers are regions of DNA sequences that, when bound by item transcription factors, can silence expression of the cistron.
Regulation by RNA [edit]
RNA can be an of import regulator of gene activity, due east.chiliad. by microRNA (miRNA), antisense-RNA, or long non-coding RNA (lncRNA). LncRNAs differ from mRNAs in the sense that they accept specified subcellular locations and functions. They were first discovered to exist located in the nucleus and chromatin, and the localizations and functions are highly diverse at present. Some yet reside in chromatin where they interact with proteins. While this lncRNA ultimately affects cistron expression in neuronal disorders such as Parkinson, Huntington, and Alzheimer disease, others, such equally, PNCTR(pyrimidine-rich non-coding transcriptors), play a part in lung cancer. Given their office in affliction, lncRNAs are potential biomarkers and may be useful targets for drugs or gene therapy, although there are no approved drugs that targert lncRNAs yet. In that location number of lncRNAs in the human genome remains poorly divers, simply some estimates range from 16,000 to 100,000 lnc genes.[v]
Epigenetic gene regulation [edit]

Overview of Epigenetic mechanisms.
Epigenetics refers to the modification of genes that is not irresolute the DNA or RNA sequence. Epigenetic modifications are also a central gene in influencing factor expression. They occur on genomic Dna and histones and their chemic modifications regulate factor expression in a more than efficient manner. There are several modifications of DNA (normally methylation) and more than 100 modifications of RNA in mammalian cells." Those modifications result in altered protein bounden to DNA and a change in RNA stability and translation efficiency.[6]
Special cases in human being biological science and illness [edit]
Regulation of transcription in cancer [edit]
In vertebrates, the majority of cistron promoters contain a CpG isle with numerous CpG sites.[7] When many of a gene's promoter CpG sites are methylated the gene becomes silenced.[8] Colorectal cancers typically have 3 to six driver mutations and 33 to 66 hitchhiker or passenger mutations.[nine] Nevertheless, transcriptional silencing may be of more importance than mutation in causing progression to cancer. For instance, in colorectal cancers nearly 600 to 800 genes are transcriptionally silenced past CpG isle methylation (see regulation of transcription in cancer). Transcriptional repression in cancer can besides occur by other epigenetic mechanisms, such as contradistinct expression of microRNAs.[x] In breast cancer, transcriptional repression of BRCA1 may occur more often by over-expressed microRNA-182 than by hypermethylation of the BRCA1 promoter (meet Low expression of BRCA1 in chest and ovarian cancers).
Regulation of transcription in addiction [edit]
One of the key features of habit is its persistence. The persistent behavioral changes appear to exist due to long-lasting changes, resulting from epigenetic alterations affecting gene expression, inside particular regions of the encephalon.[11] Drugs of abuse cause three types of epigenetic alteration in the brain. These are (1) histone acetylations and histone methylations, (2) DNA methylation at CpG sites, and (3) epigenetic downregulation or upregulation of microRNAs.[11] [12] (Meet Epigenetics of cocaine habit for some details.)
Chronic nicotine intake in mice alters brain cell epigenetic control of factor expression through acetylation of histones. This increases expression in the brain of the poly peptide FosB, important in addiction.[13] Cigarette addiction was also studied in virtually xvi,000 humans, including never smokers, current smokers, and those who had quit smoking for upwards to thirty years.[14] In blood cells, more than xviii,000 CpG sites (of the roughly 450,000 analyzed CpG sites in the genome) had frequently contradistinct methylation among current smokers. These CpG sites occurred in over 7,000 genes, or roughly a tertiary of known man genes. The majority of the differentially methylated CpG sites returned to the level of never-smokers within five years of smoking cessation. However, 2,568 CpGs among 942 genes remained differentially methylated in former versus never smokers. Such remaining epigenetic changes tin can exist viewed as "molecular scars"[12] that may affect factor expression.
In rodent models, drugs of corruption, including cocaine,[15] methamphetamine,[16] [17] alcohol[18] and tobacco smoke products,[19] all cause DNA impairment in the encephalon. During repair of DNA damages some individual repair events tin alter the methylation of DNA and/or the acetylations or methylations of histones at the sites of impairment, and thus can contribute to leaving an epigenetic scar on chromatin.[20]
Such epigenetic scars likely contribute to the persistent epigenetic changes found in addiction.
Regulation of transcription in learning and memory [edit]

DNA methylation is the add-on of a methyl group to the Deoxyribonucleic acid that happens at cytosine. The image shows a cytosine single band base of operations and a methyl group added on to the five carbon. In mammals, DNA methylation occurs nigh exclusively at a cytosine that is followed by a guanine.
In mammals, methylation of cytosine (meet Figure) in Deoxyribonucleic acid is a major regulatory mediator. Methylated cytosines primarily occur in dinucleotide sequences where cytosine is followed by a guanine, a CpG site. The full number of CpG sites in the human genome is approximately 28 meg.[21] and generally well-nigh seventy% of all CpG sites take a methylated cytosine.[22]
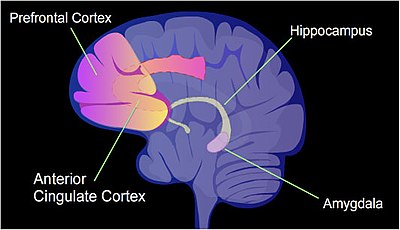
The identified areas of the human brain are involved in memory germination.
In a rat, a painful learning experience, contextual fear workout, can effect in a life-long fearful memory after a single grooming outcome.[23] Cytosine methylation is contradistinct in the promoter regions of nearly 9.17% of all genes in the hippocampus neuron DNA of a rat that has been subjected to a cursory fear conditioning experience.[24] The hippocampus is where new memories are initially stored.
Methylation of CpGs in a promoter region of a gene represses transcription[25] while methylation of CpGs in the body of a gene increases expression.[26] TET enzymes play a primal role in demethylation of methylated cytosines. Demethylation of CpGs in a cistron promoter past TET enzyme activity increases transcription of the gene.[27]
When contextual fear conditioning is practical to a rat, more than 5,000 differentially methylated regions (DMRs) (of 500 nucleotides each) occur in the rat hippocampus neural genome both one hour and 24 hours after the conditioning in the hippocampus.[24] This causes about 500 genes to be up-regulated (often due to demethylation of CpG sites in a promoter region) and nigh 1,000 genes to be down-regulated (ofttimes due to newly formed v-methylcytosine at CpG sites in a promoter region). The pattern of induced and repressed genes inside neurons appears to provide a molecular ground for forming the commencement transient retentiveness of this training event in the hippocampus of the rat brain.[24]
Post-transcriptional regulation [edit]
After the Dna is transcribed and mRNA is formed, at that place must exist some sort of regulation on how much the mRNA is translated into proteins. Cells exercise this by modulating the capping, splicing, addition of a Poly(A) Tail, the sequence-specific nuclear export rates, and, in several contexts, sequestration of the RNA transcript. These processes occur in eukaryotes simply not in prokaryotes. This modulation is a result of a protein or transcript that, in turn, is regulated and may have an affinity for sure sequences.
Three prime number untranslated regions and microRNAs [edit]
Three prime number untranslated regions (3'-UTRs) of messenger RNAs (mRNAs) often comprise regulatory sequences that mail service-transcriptionally influence factor expression.[28] Such 3'-UTRs often contain both binding sites for microRNAs (miRNAs) likewise as for regulatory proteins. By binding to specific sites within the 3'-UTR, miRNAs can subtract gene expression of various mRNAs by either inhibiting translation or direct causing degradation of the transcript. The 3'-UTR also may accept silencer regions that bind repressor proteins that inhibit the expression of a mRNA.
The 3'-UTR ofttimes contains miRNA response elements (MREs). MREs are sequences to which miRNAs bind. These are prevalent motifs within 3'-UTRs. Amongst all regulatory motifs within the 3'-UTRs (e.m. including silencer regions), MREs make up well-nigh half of the motifs.
As of 2014, the miRBase spider web site,[29] an archive of miRNA sequences and annotations, listed 28,645 entries in 233 biologic species. Of these, ane,881 miRNAs were in annotated human miRNA loci. miRNAs were predicted to have an average of about four hundred target mRNAs (affecting expression of several hundred genes).[30] Freidman et al.[30] guess that >45,000 miRNA target sites within man mRNA 3'-UTRs are conserved above background levels, and >60% of homo protein-coding genes have been nether selective pressure to maintain pairing to miRNAs.
Direct experiments show that a single miRNA can reduce the stability of hundreds of unique mRNAs.[31] Other experiments show that a single miRNA may repress the production of hundreds of proteins, but that this repression often is relatively mild (less than 2-fold).[32] [33]
The effects of miRNA dysregulation of gene expression seem to be important in cancer.[34] For instance, in gastrointestinal cancers, a 2015 newspaper identified nine miRNAs as epigenetically altered and effective in downwards-regulating Deoxyribonucleic acid repair enzymes.[35]
The effects of miRNA dysregulation of factor expression as well seem to be important in neuropsychiatric disorders, such as schizophrenia, bipolar disorder, major depressive disorder, Parkinson'south disease, Alzheimer's disease and autism spectrum disorders.[36] [37] [38]
Regulation of translation [edit]
The translation of mRNA can too be controlled by a number of mechanisms, mostly at the level of initiation. Recruitment of the pocket-size ribosomal subunit can indeed exist modulated by mRNA secondary construction, antisense RNA binding, or protein binding. In both prokaryotes and eukaryotes, a large number of RNA binding proteins exist, which ofttimes are directed to their target sequence by the secondary structure of the transcript, which may modify depending on sure atmospheric condition, such every bit temperature or presence of a ligand (aptamer). Some transcripts human activity as ribozymes and self-regulate their expression.
Examples of cistron regulation [edit]
- Enzyme induction is a process in which a molecule (eastward.g., a drug) induces (i.e., initiates or enhances) the expression of an enzyme.
- The induction of heat daze proteins in the fruit fly Drosophila melanogaster.
- The Lac operon is an interesting example of how gene expression tin be regulated.
- Viruses, despite having simply a few genes, possess mechanisms to regulate their gene expression, typically into an early and late stage, using collinear systems regulated by anti-terminators (lambda phage) or splicing modulators (HIV).
- Gal4 is a transcriptional activator that controls the expression of GAL1, GAL7, and GAL10 (all of which code for the metabolic of galactose in yeast). The GAL4/UAS arrangement has been used in a diverseness of organisms across various phyla to study gene expression.[39]
Developmental biology [edit]
A large number of studied regulatory systems come from developmental biology. Examples include:
- The colinearity of the Hox gene cluster with their nested antero-posterior patterning
- Design generation of the hand (digits - interdigits): the slope of sonic hedgehog (secreted inducing factor) from the zone of polarizing activity in the limb, which creates a gradient of active Gli3, which activates Gremlin, which inhibits BMPs also secreted in the limb, results in the germination of an alternating pattern of activity as a effect of this reaction–improvidence organization.
- Somitogenesis is the creation of segments (somites) from a uniform tissue (Pre-somitic Mesoderm). They are formed sequentially from anterior to posterior. This is accomplished in amniotes possibly by means of 2 opposing gradients, Retinoic acid in the anterior (wavefront) and Wnt and Fgf in the posterior, coupled to an aquiver blueprint (sectionalisation clock) composed of FGF + Notch and Wnt in antiphase.[twoscore]
- Sex conclusion in the soma of a Drosophila requires the sensing of the ratio of autosomal genes to sex activity chromosome-encoded genes, which results in the production of sexless splicing cistron in females, resulting in the female isoform of doublesex.[41]
Circuitry [edit]
Up-regulation and downwardly-regulation [edit]
Up-regulation is a procedure that occurs within a cell triggered by a signal (originating internal or external to the cell), which results in increased expression of one or more genes and as a effect the proteins encoded past those genes. Conversely, downward-regulation is a process resulting in decreased cistron and corresponding protein expression.
- Up-regulation occurs, for case, when a prison cell is deficient in some kind of receptor. In this case, more receptor protein is synthesized and transported to the membrane of the cell and, thus, the sensitivity of the cell is brought dorsum to normal, reestablishing homeostasis.
- Down-regulation occurs, for example, when a jail cell is overstimulated by a neurotransmitter, hormone, or drug for a prolonged menstruum of fourth dimension, and the expression of the receptor protein is decreased in guild to protect the cell (see also tachyphylaxis).
Inducible vs. repressible systems [edit]

Gene regulation works using operators and repressors in leaner.
Gene Regulation can be summarized past the response of the respective system:
- Inducible systems - An inducible system is off unless there is the presence of some molecule (chosen an inducer) that allows for gene expression. The molecule is said to "induce expression". The mode by which this happens is dependent on the control mechanisms besides equally differences betwixt prokaryotic and eukaryotic cells.
- Repressible systems - A repressible organisation is on except in the presence of some molecule (chosen a corepressor) that suppresses factor expression. The molecule is said to "repress expression". The manner past which this happens is dependent on the control mechanisms as well equally differences between prokaryotic and eukaryotic cells.
The GAL4/UAS arrangement is an example of both an inducible and repressible system. Gal4 binds an upstream activation sequence (UAS) to actuate the transcription of the GAL1/GAL7/GAL10 cassette. On the other manus, a MIG1 response to the presence of glucose can inhibit GAL4 and therefore end the expression of the GAL1/GAL7/GAL10 cassette.[42]
Theoretical circuits [edit]
- Repressor/Inducer: an activation of a sensor results in the alter of expression of a gene
- negative feedback: the factor product downregulates its own production directly or indirectly, which can result in
- keeping transcript levels constant/proportional to a factor
- inhibition of run-away reactions when coupled with a positive feedback loop
- creating an oscillator past taking advantage in the time delay of transcription and translation, given that the mRNA and protein half-life is shorter
- positive feedback: the cistron product upregulates its own product straight or indirectly, which can result in
- signal distension
- bistable switches when two genes inhibit each other and both take positive feedback
- pattern generation
Report methods [edit]
In full general, almost experiments investigating differential expression used whole cell extracts of RNA, called steady-state levels, to determine which genes inverse and by how much. These are, however, non informative of where the regulation has occurred and may mask conflicting regulatory processes (see post-transcriptional regulation), but it is still the nearly commonly analysed (quantitative PCR and Deoxyribonucleic acid microarray).
When studying cistron expression, there are several methods to look at the various stages. In eukaryotes these include:
- The local chromatin surroundings of the region can be determined by Flake-chip analysis by pulling downwardly RNA Polymerase 2, Histone 3 modifications, Trithorax-group protein, Polycomb-grouping protein, or whatsoever other DNA-bounden element to which a good antibody is available.
- Epistatic interactions can be investigated by constructed genetic array analysis
- Due to postal service-transcriptional regulation, transcription rates and total RNA levels differ significantly. To measure the transcription rates nuclear run-on assays tin be done and newer high-throughput methods are being developed, using thiol labelling instead of radioactivity.[43]
- Only 5% of the RNA polymerised in the nucleus exits,[44] and not merely introns, abortive products, and not-sense transcripts are degradated. Therefore, the differences in nuclear and cytoplasmic levels can exist encounter by separating the two fractions past gentle lysis.[45]
- Alternative splicing can exist analysed with a splicing array or with a tiling assortment (run across DNA microarray).
- All in vivo RNA is complexed equally RNPs. The quantity of transcripts bound to specific protein can be as well analysed past RIP-Chip. For instance, DCP2 will give an indication of sequestered protein; ribosome-bound gives and indication of transcripts active in transcription (although a more dated method, called polysome fractionation, is still pop in some labs)
- Protein levels can be analysed by Mass spectrometry, which can be compared only to quantitative PCR data, as microarray data is relative and non absolute.
- RNA and protein degradation rates are measured by means of transcription inhibitors (actinomycin D or α-amanitin) or translation inhibitors (Cycloheximide), respectively.
See too [edit]
- Artificial transcription factors (small molecules that mimic transcription factor protein)
- Cellular model
- Conserved non-coding Dna sequence
- Enhancer (genetics)
- Gene structure
- Spatiotemporal factor expression
Notes and references [edit]
- ^ "Can genes be turned on and off in cells?". Genetics Home Reference.
- ^ Bell JT, Pai AA, Pickrell JK, Gaffney DJ, Pique-Regi R, Degner JF, et al. (2011). "DNA methylation patterns associate with genetic and gene expression variation in HapMap cell lines". Genome Biology. 12 (1): R10. doi:10.1186/gb-2011-12-1-r10. PMC3091299. PMID 21251332.
- ^ Vertino PM, Spillare EA, Harris CC, Baylin SB (April 1993). "Altered chromosomal methylation patterns back-trail oncogene-induced transformation of human bronchial epithelial cells" (PDF). Cancer Research. 53 (7): 1684–9. PMID 8453642.
- ^ Austin S, Dixon R (June 1992). "The prokaryotic enhancer binding protein NTRC has an ATPase activity which is phosphorylation and Deoxyribonucleic acid dependent". The EMBO Periodical. xi (vi): 2219–28. doi:10.1002/j.1460-2075.1992.tb05281.x. PMC556689. PMID 1534752.
- ^ Statello L, Guo CJ, Chen LL, Huarte Thousand (February 2021). "Gene regulation by long non-coding RNAs and its biological functions". Nature Reviews. Molecular Cell Biology. 22 (ii): 96–118. doi:ten.1038/s41580-020-00315-9. ISSN 1471-0072. PMC7754182. PMID 33353982.
- ^ Kan RL, Chen J, Sallam T (July 2021). "Crosstalk between epitranscriptomic and epigenetic mechanisms in gene regulation". Trends in Genetics. 38 (2): 182–193. doi:ten.1016/j.tig.2021.06.014. PMID 34294427. S2CID 236200223.
- ^ Saxonov South, Berg P, Brutlag DL (January 2006). "A genome-wide analysis of CpG dinucleotides in the human genome distinguishes two distinct classes of promoters". Proceedings of the National Academy of Sciences of the United states of America. 103 (5): 1412–vii. Bibcode:2006PNAS..103.1412S. doi:10.1073/pnas.0510310103. PMC1345710. PMID 16432200.
- ^ Bird A (January 2002). "DNA methylation patterns and epigenetic memory". Genes & Development. sixteen (i): 6–21. doi:10.1101/gad.947102. PMID 11782440.
- ^ Vogelstein B, Papadopoulos Due north, Velculescu VE, Zhou S, Diaz LA, Kinzler KW (March 2013). "Cancer genome landscapes". Science. 339 (6127): 1546–58. Bibcode:2013Sci...339.1546V. doi:ten.1126/science.1235122. PMC3749880. PMID 23539594.
- ^ Tessitore A, Cicciarelli G, Del Vecchio F, Gaggiano A, Verzella D, Fischietti M, et al. (2014). "MicroRNAs in the DNA Harm/Repair Network and Cancer". International Journal of Genomics. 2014: 820248. doi:10.1155/2014/820248. PMC3926391. PMID 24616890.
- ^ a b Nestler EJ (January 2014). "Epigenetic mechanisms of drug addiction". Neuropharmacology. 76 Pt B: 259–68. doi:10.1016/j.neuropharm.2013.04.004. PMC3766384. PMID 23643695.
- ^ a b Robison AJ, Nestler EJ (October 2011). "Transcriptional and epigenetic mechanisms of addiction". Nature Reviews. Neuroscience. 12 (11): 623–37. doi:x.1038/nrn3111. PMC3272277. PMID 21989194.
- ^ Levine A, Huang Y, Drisaldi B, Griffin EA, Pollak DD, Xu Southward, et al. (November 2011). "Molecular machinery for a gateway drug: epigenetic changes initiated by nicotine prime cistron expression by cocaine". Science Translational Medicine. 3 (107): 107ra109. doi:ten.1126/scitranslmed.3003062. PMC4042673. PMID 22049069.
- ^ Joehanes R, Just Ac, Marioni RE, Pilling LC, Reynolds LM, Mandaviya PR, et al. (Oct 2016). "Epigenetic Signatures of Cigarette Smoking". Apportionment: Cardiovascular Genetics. nine (5): 436–447. doi:10.1161/CIRCGENETICS.116.001506. PMC5267325. PMID 27651444.
- ^ de Souza MF, Gonçales TA, Steinmetz A, Moura DJ, Saffi J, Gomez R, Barros HM (April 2014). "Cocaine induces DNA damage in distinct brain areas of female rats under different hormonal conditions". Clinical and Experimental Pharmacology & Physiology. 41 (4): 265–nine. doi:ten.1111/1440-1681.12218. PMID 24552452. S2CID 20849951.
- ^ Johnson Z, Venters J, Guarraci FA, Zewail-Foote K (June 2015). "Methamphetamine induces Dna damage in specific regions of the female rat encephalon". Clinical and Experimental Pharmacology & Physiology. 42 (six): 570–v. doi:10.1111/1440-1681.12404. PMID 25867833. S2CID 24182756.
- ^ Tokunaga I, Ishigami A, Kubo S, Gotohda T, Kitamura O (August 2008). "The peroxidative Deoxyribonucleic acid damage and apoptosis in methamphetamine-treated rat brain". The Journal of Medical Investigation. 55 (3–four): 241–5. doi:10.2152/jmi.55.241. PMID 18797138.
- ^ Rulten SL, Hodder E, Ripley TL, Stephens DN, Mayne LV (July 2008). "Alcohol induces DNA damage and the Fanconi anemia D2 poly peptide implicating FANCD2 in the Dna damage response pathways in brain". Alcoholism, Clinical and Experimental Inquiry. 32 (7): 1186–96. doi:x.1111/j.1530-0277.2008.00673.x. PMID 18482162.
- ^ Adhami N, Chen Y, Martins-Green One thousand (October 2017). "Biomarkers of disease tin be detected in mice equally early on equally four weeks after initiation of exposure to third-paw smoke levels equivalent to those found in homes of smokers". Clinical Science. 131 (nineteen): 2409–2426. doi:x.1042/CS20171053. PMID 28912356.
- ^ Dabin J, Fortuny A, Polo SE (June 2016). "Epigenome Maintenance in Response to DNA Damage". Molecular Cell. 62 (five): 712–27. doi:ten.1016/j.molcel.2016.04.006. PMC5476208. PMID 27259203.
- ^ Lövkvist C, Dodd IB, Sneppen K, Haerter JO (June 2016). "Deoxyribonucleic acid methylation in human epigenomes depends on local topology of CpG sites". Nucleic Acids Research. 44 (11): 5123–32. doi:ten.1093/nar/gkw124. PMC4914085. PMID 26932361.
- ^ Jabbari K, Bernardi G (May 2004). "Cytosine methylation and CpG, TpG (CpA) and TpA frequencies". Gene. 333: 143–9. doi:10.1016/j.factor.2004.02.043. PMID 15177689.
- ^ Kim JJ, Jung MW (2006). "Neural circuits and mechanisms involved in Pavlovian fear workout: a critical review". Neuroscience and Biobehavioral Reviews. 30 (ii): 188–202. doi:ten.1016/j.neubiorev.2005.06.005. PMC4342048. PMID 16120461.
- ^ a b c Duke CG, Kennedy AJ, Gavin CF, Day JJ, Sweatt JD (July 2017). "Feel-dependent epigenomic reorganization in the hippocampus". Learning & Memory. 24 (7): 278–288. doi:10.1101/lm.045112.117. PMC5473107. PMID 28620075.
- ^ Weber M, Hellmann I, Stadler MB, Ramos L, Pääbo Due south, Rebhan M, Schübeler D (April 2007). "Distribution, silencing potential and evolutionary impact of promoter DNA methylation in the human genome". Nat. Genet. 39 (4): 457–66. doi:10.1038/ng1990. PMID 17334365. S2CID 22446734.
- ^ Yang Ten, Han H, De Carvalho DD, Lay FD, Jones PA, Liang G (Oct 2014). "Gene trunk methylation tin alter gene expression and is a therapeutic target in cancer". Cancer Cell. 26 (4): 577–90. doi:10.1016/j.ccr.2014.07.028. PMC4224113. PMID 25263941.
- ^ Maeder ML, Angstman JF, Richardson ME, Linder SJ, Cascio VM, Tsai SQ, Ho QH, Sander JD, Reyon D, Bernstein Exist, Costello JF, Wilkinson MF, Joung JK (December 2013). "Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins". Nat. Biotechnol. 31 (12): 1137–42. doi:10.1038/nbt.2726. PMC3858462. PMID 24108092.
- ^ Ogorodnikov A, Kargapolova Y, Danckwardt South (June 2016). "Processing and transcriptome expansion at the mRNA 3' end in health and disease: finding the correct stop". Pflügers Archiv. 468 (six): 993–1012. doi:x.1007/s00424-016-1828-three. PMC4893057. PMID 27220521.
- ^ miRBase.org
- ^ a b Friedman RC, Farh KK, Burge CB, Bartel DP (Jan 2009). "Most mammalian mRNAs are conserved targets of microRNAs". Genome Inquiry. 19 (1): 92–105. doi:10.1101/gr.082701.108. PMC2612969. PMID 18955434.
- ^ Lim LP, Lau NC, Garrett-Engele P, Grimson A, Schelter JM, Castle J, et al. (February 2005). "Microarray analysis shows that some microRNAs downregulate big numbers of target mRNAs". Nature. 433 (7027): 769–73. Bibcode:2005Natur.433..769L. doi:10.1038/nature03315. PMID 15685193. S2CID 4430576.
- ^ Selbach Yard, Schwanhäusser B, Thierfelder N, Fang Z, Khanin R, Rajewsky N (September 2008). "Widespread changes in protein synthesis induced by microRNAs". Nature. 455 (7209): 58–63. Bibcode:2008Natur.455...58S. doi:10.1038/nature07228. PMID 18668040. S2CID 4429008.
- ^ Baek D, Villén J, Shin C, Camargo FD, Gygi SP, Bartel DP (September 2008). "The impact of microRNAs on protein output". Nature. 455 (7209): 64–71. Bibcode:2008Natur.455...64B. doi:10.1038/nature07242. PMC2745094. PMID 18668037.
- ^ Palmero EI, de Campos SG, Campos One thousand, de Souza NC, Guerreiro ID, Carvalho AL, Marques MM (July 2011). "Mechanisms and role of microRNA deregulation in cancer onset and progression". Genetics and Molecular Biological science. 34 (3): 363–70. doi:10.1590/S1415-47572011000300001. PMC3168173. PMID 21931505.
- ^ Bernstein C, Bernstein H (May 2015). "Epigenetic reduction of Dna repair in progression to gastrointestinal cancer". Earth Journal of Gastrointestinal Oncology. 7 (5): 30–46. doi:10.4251/wjgo.v7.i5.30. PMC4434036. PMID 25987950.
- ^ Maffioletti E, Tardito D, Gennarelli M, Bocchio-Chiavetto Fifty (2014). "Micro spies from the brain to the periphery: new clues from studies on microRNAs in neuropsychiatric disorders". Frontiers in Cellular Neuroscience. viii: 75. doi:10.3389/fncel.2014.00075. PMC3949217. PMID 24653674.
- ^ Mellios North, Sur M (2012). "The Emerging Role of microRNAs in Schizophrenia and Autism Spectrum Disorders". Frontiers in Psychiatry. 3: 39. doi:ten.3389/fpsyt.2012.00039. PMC3336189. PMID 22539927.
- ^ Geaghan M, Cairns MJ (August 2015). "MicroRNA and Posttranscriptional Dysregulation in Psychiatry". Biological Psychiatry. 78 (4): 231–9. doi:x.1016/j.biopsych.2014.12.009. PMID 25636176.
- ^ Barnett JA (July 2004). "A history of research on yeasts vii: enzymic accommodation and regulation". Yeast. 21 (9): 703–46. doi:x.1002/yea.1113. PMID 15282797. S2CID 36606279.
- ^ Dequéant ML, Pourquié O (May 2008). "Segmental patterning of the vertebrate embryonic axis". Nature Reviews. Genetics. 9 (v): 370–82. doi:x.1038/nrg2320. PMID 18414404. S2CID 2526914.
- ^ Gilbert SF (2003). Developmental biology, 7th ed., Sunderland, Mass: Sinauer Associates, 65–6. ISBN 0-87893-258-5.
- ^ Nehlin JO, Carlberg G, Ronne H (November 1991). "Control of yeast GAL genes by MIG1 repressor: a transcriptional cascade in the glucose response". The EMBO Periodical. 10 (11): 3373–vii. doi:10.1002/j.1460-2075.1991.tb04901.x. PMC453065. PMID 1915298.
- ^ Cheadle C, Fan J, Cho-Chung YS, Werner T, Ray J, Exercise L, et al. (May 2005). "Command of gene expression during T prison cell activation: alternating regulation of mRNA transcription and mRNA stability". BMC Genomics. 6: 75. doi:10.1186/1471-2164-6-75. PMC1156890. PMID 15907206.
- ^ Jackson DA, Pombo A, Iborra F (February 2000). "The balance sheet for transcription: an analysis of nuclear RNA metabolism in mammalian cells". FASEB Periodical. xiv (2): 242–54. doi:10.1096/fasebj.14.2.242. PMID 10657981. S2CID 23518786.
- ^ Schwanekamp JA, Sartor MA, Karyala S, Halbleib D, Medvedovic Grand, Tomlinson CR (2006). "Genome-wide analyses testify that nuclear and cytoplasmic RNA levels are differentially afflicted by dioxin". Biochimica et Biophysica Acta (BBA) - Cistron Construction and Expression. 1759 (viii–9): 388–402. doi:10.1016/j.bbaexp.2006.07.005. PMID 16962184.
Bibliography [edit]
- Latchman, David S. (2005). Gene regulation: a eukaryotic perspective. Psychology Press. ISBN978-0-415-36510-ix.
External links [edit]
- Establish Transcription Gene Database and Plant Transcriptional Regulation Data and Analysis Platform
- Regulation of Gene Expression (MeSH) at the US National Library of Medicine Medical Subject Headings (MeSH)
- ChIPBase An open database for decoding the transcriptional regulatory networks of non-coding RNAs and protein-coding genes from Bit-seq data.
Source: https://en.wikipedia.org/wiki/Regulation_of_gene_expression
Mag-post ng isang Komento for "Drag the Terms to Their Corresponding Class in Order to Review Various Aspects of Gene Regulation"