What is the term used to describe the scientific study of heredity?
Genetics is a branch of biological science concerned with the report of genes, genetic variation, and heredity in organisms.[one] [2] [three]
Though heredity had been observed for millennia, Gregor Mendel, Moravian scientist and Augustinian friar working in the 19th century in Brno, was the first to study genetics scientifically. Mendel studied "trait inheritance", patterns in the mode traits are handed down from parents to offspring over fourth dimension. He observed that organisms (pea plants) inherit traits by way of discrete "units of inheritance". This term, withal used today, is a somewhat cryptic definition of what is referred to equally a cistron.
Trait inheritance and molecular inheritance mechanisms of genes are yet master principles of genetics in the 21st century, but mod genetics has expanded beyond inheritance to studying the function and behavior of genes. Gene construction and part, variation, and distribution are studied within the context of the cell, the organism (e.yard. dominance), and within the context of a population. Genetics has given rising to a number of subfields, including molecular genetics, epigenetics and population genetics. Organisms studied within the broad field span the domains of life (archaea, bacteria, and eukarya).
Genetic processes work in combination with an organism'southward environment and experiences to influence development and beliefs, often referred to every bit nature versus nurture. The intracellular or extracellular environment of a living cell or organism may switch factor transcription on or off. A classic example is ii seeds of genetically identical corn, one placed in a temperate climate and one in an barren climate (lacking sufficient waterfall or rain). While the average height of the 2 corn stalks may exist genetically adamant to be equal, the ane in the arid climate just grows to half the height of the one in the temperate climate due to lack of water and nutrients in its environment.
Etymology [edit]
The word genetics stems from the ancient Greek γενετικός genetikos pregnant "genitive"/"generative", which in turn derives from γένεσις genesis significant "origin".[4] [5] [6]
History [edit]
The observation that living things inherit traits from their parents has been used since prehistoric times to better ingather plants and animals through selective convenance.[seven] The modern science of genetics, seeking to empathize this process, began with the piece of work of the Augustinian friar Gregor Mendel in the mid-19th century.[8]
Prior to Mendel, Imre Festetics, a Hungarian noble, who lived in Kőszeg before Mendel, was the first who used the give-and-take "genetics." He described several rules of genetic inheritance in his piece of work The genetic law of the Nature (Die genetischen Gesetze der Natur, 1819). His second police force is the same every bit what Mendel published. In his tertiary law, he developed the basic principles of mutation (he tin be considered a forerunner of Hugo de Vries).[9]

Other theories of inheritance preceded Mendel'southward work. A popular theory during the 19th century, and implied by Charles Darwin's 1859 On the Origin of Species, was blending inheritance: the thought that individuals inherit a smooth blend of traits from their parents.[10] Mendel'southward work provided examples where traits were definitely non blended after hybridization, showing that traits are produced past combinations of distinct genes rather than a continuous alloy. Blending of traits in the progeny is now explained by the activeness of multiple genes with quantitative effects. Another theory that had some back up at that time was the inheritance of acquired characteristics: the conventionalities that individuals inherit traits strengthened by their parents. This theory (usually associated with Jean-Baptiste Lamarck) is now known to exist wrong—the experiences of individuals do non impact the genes they pass to their children,[xi] Other theories included the pangenesis of Charles Darwin (which had both acquired and inherited aspects) and Francis Galton's reformulation of pangenesis as both particulate and inherited.[12]
Mendelian and classical genetics [edit]

Morgan'southward observation of sex-linked inheritance of a mutation causing white eyes in Drosophila led him to the hypothesis that genes are located upon chromosomes.
Modern genetics started with Mendel'due south studies of the nature of inheritance in plants. In his paper "Versuche über Pflanzenhybriden" ("Experiments on Constitute Hybridization"), presented in 1865 to the Naturforschender Verein (Social club for Enquiry in Nature) in Brünn, Mendel traced the inheritance patterns of certain traits in pea plants and described them mathematically.[xiii] Although this blueprint of inheritance could just be observed for a few traits, Mendel'due south work suggested that heredity was particulate, not acquired, and that the inheritance patterns of many traits could exist explained through simple rules and ratios.
The importance of Mendel's work did not gain wide understanding until 1900, afterwards his death, when Hugo de Vries and other scientists rediscovered his research. William Bateson, a proponent of Mendel's work, coined the word genetics in 1905[14] [15] (the adjective genetic, derived from the Greek word genesis—γένεσις, "origin", predates the noun and was showtime used in a biological sense in 1860[16]). Bateson both acted equally a mentor and was aided significantly by the work of other scientists from Newnham Higher at Cambridge, specifically the piece of work of Becky Saunders, Nora Darwin Barlow, and Muriel Wheldale Onslow.[17] Bateson popularized the usage of the word genetics to depict the study of inheritance in his inaugural address to the Tertiary International Conference on Plant Hybridization in London in 1906.[eighteen]
After the rediscovery of Mendel's work, scientists tried to decide which molecules in the jail cell were responsible for inheritance. In 1900, Nettie Stevens began studying the mealworm.[nineteen] Over the adjacent xi years, she discovered that females only had the Ten chromosome and males had both X and Y chromosomes.[nineteen] She was able to conclude that sex is a chromosomal factor and is determined past the male.[nineteen] In 1911, Thomas Hunt Morgan argued that genes are on chromosomes, based on observations of a sex-linked white centre mutation in fruit flies.[twenty] In 1913, his educatee Alfred Sturtevant used the phenomenon of genetic linkage to show that genes are bundled linearly on the chromosome.[21]
Molecular genetics [edit]

Deoxyribonucleic acid, the molecular basis for biological inheritance. Each strand of DNA is a chain of nucleotides, matching each other in the center to course what look like rungs on a twisted ladder.
Although genes were known to exist on chromosomes, chromosomes are composed of both poly peptide and Dna, and scientists did not know which of the two is responsible for inheritance. In 1928, Frederick Griffith discovered the phenomenon of transformation (see Griffith's experiment): dead bacteria could transfer genetic material to "transform" other still-living bacteria. Sixteen years subsequently, in 1944, the Avery–MacLeod–McCarty experiment identified Dna equally the molecule responsible for transformation.[22] The function of the nucleus as the repository of genetic information in eukaryotes had been established by Hämmerling in 1943 in his work on the single celled alga Acetabularia.[23] The Hershey–Hunt experiment in 1952 confirmed that Deoxyribonucleic acid (rather than poly peptide) is the genetic cloth of the viruses that infect bacteria, providing farther evidence that Deoxyribonucleic acid is the molecule responsible for inheritance.[24]
James Watson and Francis Crick adamant the structure of Deoxyribonucleic acid in 1953, using the 10-ray crystallography work of Rosalind Franklin and Maurice Wilkins that indicated DNA has a helical construction (i.e., shaped similar a corkscrew).[25] [26] Their double-helix model had two strands of DNA with the nucleotides pointing inwards, each matching a complementary nucleotide on the other strand to form what await like rungs on a twisted ladder.[27] This structure showed that genetic information exists in the sequence of nucleotides on each strand of Deoxyribonucleic acid. The construction also suggested a simple method for replication: if the strands are separated, new partner strands tin can be reconstructed for each based on the sequence of the old strand. This property is what gives DNA its semi-conservative nature where i strand of new Dna is from an original parent strand.[28]
Although the structure of DNA showed how inheritance works, it was still not known how DNA influences the behavior of cells. In the post-obit years, scientists tried to understand how DNA controls the process of poly peptide product.[29] It was discovered that the cell uses DNA as a template to create matching messenger RNA, molecules with nucleotides very like to Deoxyribonucleic acid. The nucleotide sequence of a messenger RNA is used to create an amino acid sequence in poly peptide; this translation betwixt nucleotide sequences and amino acid sequences is known as the genetic code.[xxx]
With the newfound molecular agreement of inheritance came an explosion of enquiry.[31] A notable theory arose from Tomoko Ohta in 1973 with her amendment to the neutral theory of molecular development through publishing the nearly neutral theory of molecular evolution. In this theory, Ohta stressed the importance of natural selection and the environment to the rate at which genetic development occurs.[32] 1 important development was concatenation-termination DNA sequencing in 1977 by Frederick Sanger. This technology allows scientists to read the nucleotide sequence of a DNA molecule.[33] In 1983, Kary Banks Mullis adult the polymerase chain reaction, providing a quick way to isolate and amplify a specific department of Dna from a mixture.[34] The efforts of the Human Genome Projection, Section of Energy, NIH, and parallel private efforts by Celera Genomics led to the sequencing of the human genome in 2003.[35] [36]
Features of inheritance [edit]
Discrete inheritance and Mendel's laws [edit]
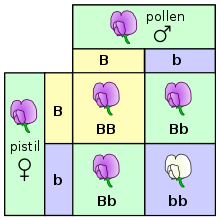
A Punnett square depicting a cross betwixt two pea plants heterozygous for majestic (B) and white (b) blossoms.
At its most fundamental level, inheritance in organisms occurs by passing discrete heritable units, called genes, from parents to offspring.[37] This property was get-go observed by Gregor Mendel, who studied the segregation of heritable traits in pea plants.[13] [38] In his experiments studying the trait for blossom colour, Mendel observed that the flowers of each pea establish were either royal or white—simply never an intermediate betwixt the two colors. These different, discrete versions of the same gene are chosen alleles.
In the case of the pea, which is a diploid species, each private plant has two copies of each gene, one copy inherited from each parent.[39] Many species, including humans, accept this blueprint of inheritance. Diploid organisms with two copies of the same allele of a given gene are called homozygous at that gene locus, while organisms with two different alleles of a given gene are called heterozygous.
The set of alleles for a given organism is called its genotype, while the observable traits of the organism are chosen its phenotype. When organisms are heterozygous at a gene, oft one allele is called dominant as its qualities dominate the phenotype of the organism, while the other allele is chosen recessive as its qualities recede and are not observed. Some alleles do not accept complete authorisation and instead have incomplete authorisation by expressing an intermediate phenotype, or codominance by expressing both alleles at once.[xl]
When a pair of organisms reproduce sexually, their offspring randomly inherit one of the two alleles from each parent. These observations of discrete inheritance and the segregation of alleles are collectively known as Mendel'southward commencement law or the Law of Segregation.
Notation and diagrams [edit]

Genetic pedigree charts help rails the inheritance patterns of traits.
Geneticists use diagrams and symbols to draw inheritance. A gene is represented by one or a few letters. Often a "+" symbol is used to marker the usual, non-mutant allele for a gene.[41]
In fertilization and convenance experiments (and especially when discussing Mendel's laws) the parents are referred to as the "P" generation and the offspring as the "F1" (beginning filial) generation. When the F1 offspring mate with each other, the offspring are chosen the "F2" (2nd filial) generation. One of the common diagrams used to predict the result of cross-breeding is the Punnett square.
When studying human genetic diseases, geneticists oft use pedigree charts to correspond the inheritance of traits.[42] These charts map the inheritance of a trait in a family unit tree.
Multiple factor interactions [edit]
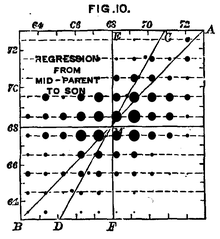
Human height is a trait with complex genetic causes. Francis Galton'due south information from 1889 shows the relationship betwixt offspring pinnacle as a function of mean parent height.
Organisms have thousands of genes, and in sexually reproducing organisms these genes generally assort independently of each other. This ways that the inheritance of an allele for yellow or green pea colour is unrelated to the inheritance of alleles for white or purple flowers. This phenomenon, known as "Mendel's second law" or the "law of contained assortment," means that the alleles of different genes get shuffled betwixt parents to form offspring with many different combinations. (Some genes do not assort independently, demonstrating genetic linkage, a topic discussed later in this article.)
Frequently different genes tin collaborate in a way that influences the same trait. In the Blue-eyed Mary (Omphalodes verna), for case, at that place exists a gene with alleles that determine the color of flowers: blue or magenta. Another gene, however, controls whether the flowers have color at all or are white. When a plant has 2 copies of this white allele, its flowers are white—regardless of whether the first gene has blue or magenta alleles. This interaction between genes is called epistasis, with the second factor epistatic to the first.[43]
Many traits are not discrete features (e.1000. purple or white flowers) but are instead continuous features (e.g. human top and skin color). These complex traits are products of many genes.[44] The influence of these genes is mediated, to varying degrees, past the environment an organism has experienced. The degree to which an organism'southward genes contribute to a complex trait is called heritability.[45] Measurement of the heritability of a trait is relative—in a more variable environment, the environment has a bigger influence on the total variation of the trait. For example, human superlative is a trait with complex causes. Information technology has a heritability of 89% in the United states. In Nigeria, however, where people experience a more variable access to good nutrition and health care, superlative has a heritability of just 62%.[46]
Molecular basis for inheritance [edit]
Dna and chromosomes [edit]


The molecular basis for genes is deoxyribonucleic acid (Dna). Deoxyribonucleic acid is equanimous of a concatenation of nucleotides, of which at that place are four types: adenine (A), cytosine (C), guanine (M), and thymine (T). Genetic information exists in the sequence of these nucleotides, and genes be equally stretches of sequence along the Deoxyribonucleic acid chain.[47] Viruses are the only exception to this rule—sometimes viruses use the very like molecule RNA instead of Deoxyribonucleic acid as their genetic fabric.[48] Viruses cannot reproduce without a host and are unaffected by many genetic processes, then tend not to be considered living organisms.
Deoxyribonucleic acid ordinarily exists as a double-stranded molecule, coiled into the shape of a double helix. Each nucleotide in Deoxyribonucleic acid preferentially pairs with its partner nucleotide on the contrary strand: A pairs with T, and C pairs with M. Thus, in its two-stranded grade, each strand effectively contains all necessary data, redundant with its partner strand. This construction of DNA is the physical basis for inheritance: DNA replication duplicates the genetic information by splitting the strands and using each strand as a template for synthesis of a new partner strand.[49]
Genes are arranged linearly along long chains of DNA base-pair sequences. In leaner, each cell normally contains a unmarried round genophore, while eukaryotic organisms (such equally plants and animals) have their DNA arranged in multiple linear chromosomes. These DNA strands are often extremely long; the largest man chromosome, for example, is nigh 247 million base pairs in length.[50] The DNA of a chromosome is associated with structural proteins that organize, compact, and control access to the Deoxyribonucleic acid, forming a material called chromatin; in eukaryotes, chromatin is usually equanimous of nucleosomes, segments of Deoxyribonucleic acid wound effectually cores of histone proteins.[51] The full fix of hereditary material in an organism (usually the combined DNA sequences of all chromosomes) is chosen the genome.
DNA is most often establish in the nucleus of cells, but Ruth Sager helped in the discovery of nonchromosomal genes establish outside of the nucleus.[52] In plants, these are often institute in the chloroplasts and in other organisms, in the mitochondria.[52] These nonchromosomal genes tin can still be passed on by either partner in sexual reproduction and they control a variety of hereditary characteristics that replicate and remain active throughout generations.[52]
While haploid organisms have only one copy of each chromosome, about animals and many plants are diploid, containing two of each chromosome and thus two copies of every gene.[39] The two alleles for a cistron are located on identical loci of the ii homologous chromosomes, each allele inherited from a different parent.

Walther Flemming's 1882 diagram of eukaryotic prison cell division. Chromosomes are copied, condensed, and organized. And then, equally the cell divides, chromosome copies carve up into the daughter cells.
Many species have and then-called sex chromosomes that decide the gender of each organism.[53] In humans and many other animals, the Y chromosome contains the factor that triggers the development of the specifically male person characteristics. In development, this chromosome has lost almost of its content and also about of its genes, while the X chromosome is similar to the other chromosomes and contains many genes. This being said, Mary Frances Lyon discovered that there is X-chromosome inactivation during reproduction to avert passing on twice as many genes to the offspring.[54] Lyon's discovery led to the discovery of other things including X-linked diseases.[54] The 10 and Y chromosomes form a strongly heterogeneous pair.
Reproduction [edit]
When cells divide, their full genome is copied and each daughter jail cell inherits one copy. This process, called mitosis, is the simplest course of reproduction and is the basis for asexual reproduction. Asexual reproduction can besides occur in multicellular organisms, producing offspring that inherit their genome from a single parent. Offspring that are genetically identical to their parents are called clones.
Eukaryotic organisms oftentimes use sexual reproduction to generate offspring that contain a mixture of genetic material inherited from two different parents. The process of sexual reproduction alternates between forms that incorporate single copies of the genome (haploid) and double copies (diploid).[39] Haploid cells fuse and combine genetic fabric to create a diploid cell with paired chromosomes. Diploid organisms course haploids past dividing, without replicating their DNA, to create daughter cells that randomly inherit one of each pair of chromosomes. Most animals and many plants are diploid for most of their lifespan, with the haploid form reduced to single cell gametes such as sperm or eggs.
Although they do non use the haploid/diploid method of sexual reproduction, bacteria have many methods of acquiring new genetic information. Some bacteria can undergo conjugation, transferring a pocket-sized circular piece of Dna to another bacterium.[55] Leaner can besides take up raw DNA fragments found in the environment and integrate them into their genomes, a phenomenon known as transformation.[56] These processes result in horizontal gene transfer, transmitting fragments of genetic data betwixt organisms that would be otherwise unrelated. Natural bacterial transformation occurs in many bacterial species, and can exist regarded as a sexual process for transferring DNA from ane cell to another cell (usually of the aforementioned species).[57] Transformation requires the action of numerous bacterial gene products, and its master adaptive function appears to be repair of Deoxyribonucleic acid damages in the recipient cell.[57]
Recombination and genetic linkage [edit]

The diploid nature of chromosomes allows for genes on different chromosomes to assort independently or be separated from their homologous pair during sexual reproduction wherein haploid gametes are formed. In this manner new combinations of genes tin occur in the offspring of a mating pair. Genes on the same chromosome would theoretically never recombine. However, they do, via the cellular process of chromosomal crossover. During crossover, chromosomes commutation stretches of Dna, effectively shuffling the gene alleles between the chromosomes.[58] This process of chromosomal crossover generally occurs during meiosis, a series of jail cell divisions that creates haploid cells. Meiotic recombination, specially in microbial eukaryotes, appears to serve the adaptive part of repair of DNA damages.[57]
The first cytological demonstration of crossing over was performed by Harriet Creighton and Barbara McClintock in 1931. Their inquiry and experiments on corn provided cytological evidence for the genetic theory that linked genes on paired chromosomes do in fact substitution places from one homolog to the other.[59]
The probability of chromosomal crossover occurring between two given points on the chromosome is related to the distance between the points. For an arbitrarily long distance, the probability of crossover is high enough that the inheritance of the genes is effectively uncorrelated.[60] For genes that are closer together, all the same, the lower probability of crossover ways that the genes demonstrate genetic linkage; alleles for the 2 genes tend to be inherited together. The amounts of linkage between a series of genes tin can exist combined to grade a linear linkage map that roughly describes the arrangement of the genes along the chromosome.[61]
Gene expression [edit]
Genetic lawmaking [edit]

Genes by and large limited their functional effect through the production of proteins, which are complex molecules responsible for nearly functions in the cell. Proteins are made up of one or more polypeptide chains, each of which is composed of a sequence of amino acids, and the Deoxyribonucleic acid sequence of a gene (through an RNA intermediate) is used to produce a specific amino acid sequence. This process begins with the product of an RNA molecule with a sequence matching the gene's DNA sequence, a procedure called transcription.
This messenger RNA molecule and so serves to produce a corresponding amino acid sequence through a process chosen translation. Each group of three nucleotides in the sequence, chosen a codon, corresponds either to one of the twenty possible amino acids in a protein or an instruction to end the amino acid sequence; this correspondence is called the genetic code.[62] The flow of information is unidirectional: data is transferred from nucleotide sequences into the amino acid sequence of proteins, but information technology never transfers from protein back into the sequence of DNA—a phenomenon Francis Crick chosen the fundamental dogma of molecular biology.[63]
The specific sequence of amino acids results in a unique iii-dimensional structure for that poly peptide, and the three-dimensional structures of proteins are related to their functions.[64] [65] Some are simple structural molecules, like the fibers formed by the poly peptide collagen. Proteins can bind to other proteins and simple molecules, sometimes acting every bit enzymes by facilitating chemic reactions inside the jump molecules (without changing the structure of the protein itself). Protein structure is dynamic; the protein hemoglobin bends into slightly different forms equally information technology facilitates the capture, transport, and release of oxygen molecules within mammalian blood.
A single nucleotide difference inside Deoxyribonucleic acid tin can cause a change in the amino acrid sequence of a protein. Considering protein structures are the result of their amino acid sequences, some changes tin can dramatically change the properties of a protein by destabilizing the structure or changing the surface of the protein in a way that changes its interaction with other proteins and molecules. For example, sickle-cell anemia is a human genetic illness that results from a unmarried base difference within the coding region for the β-globin department of hemoglobin, causing a unmarried amino acid modify that changes hemoglobin's physical backdrop.[66] Sickle-prison cell versions of hemoglobin stick to themselves, stacking to form fibers that misconstrue the shape of red blood cells carrying the protein. These sickle-shaped cells no longer catamenia smoothly through claret vessels, having a trend to clog or degrade, causing the medical problems associated with this disease.
Some DNA sequences are transcribed into RNA but are not translated into protein products—such RNA molecules are chosen non-coding RNA. In some cases, these products fold into structures which are involved in critical prison cell functions (e.g. ribosomal RNA and transfer RNA). RNA can also have regulatory furnishings through hybridization interactions with other RNA molecules (such as microRNA).
Nature and nurture [edit]

Siamese cats have a temperature-sensitive pigment-production mutation.
Although genes contain all the information an organism uses to function, the environment plays an of import role in determining the ultimate phenotypes an organism displays. The phrase "nature and nurture" refers to this complementary human relationship. The phenotype of an organism depends on the interaction of genes and the environment. An interesting example is the coat coloration of the Siamese cat. In this case, the body temperature of the cat plays the part of the surround. The true cat's genes code for dark pilus, thus the hair-producing cells in the cat brand cellular proteins resulting in dark hair. But these nighttime hair-producing proteins are sensitive to temperature (i.eastward. accept a mutation causing temperature-sensitivity) and denature in college-temperature environments, failing to produce dark-hair pigment in areas where the cat has a college body temperature. In a low-temperature environment, all the same, the protein's structure is stable and produces nighttime-hair pigment ordinarily. The protein remains functional in areas of peel that are colder—such as its legs, ears, tail, and face—then the cat has dark hair at its extremities.[67]
Surround plays a major function in furnishings of the homo genetic disease phenylketonuria.[68] The mutation that causes phenylketonuria disrupts the ability of the trunk to break down the amino acid phenylalanine, causing a toxic build-upward of an intermediate molecule that, in turn, causes severe symptoms of progressive intellectual disability and seizures. Notwithstanding, if someone with the phenylketonuria mutation follows a strict diet that avoids this amino acrid, they remain normal and healthy.
A common method for determining how genes and environment ("nature and nurture") contribute to a phenotype involves studying identical and fraternal twins, or other siblings of multiple births.[69] Identical siblings are genetically the same since they come up from the aforementioned zygote. Meanwhile, congenial twins are as genetically unlike from one some other as normal siblings. By comparing how often a certain disorder occurs in a pair of identical twins to how frequently it occurs in a pair of fraternal twins, scientists can determine whether that disorder is acquired by genetic or postnatal environmental factors. One famous case involved the study of the Genain quadruplets, who were identical quadruplets all diagnosed with schizophrenia.[70] Still, such tests cannot split up genetic factors from ecology factors affecting fetal development.
Gene regulation [edit]
The genome of a given organism contains thousands of genes, just non all these genes need to be active at any given moment. A gene is expressed when it is existence transcribed into mRNA and at that place exist many cellular methods of controlling the expression of genes such that proteins are produced only when needed by the prison cell. Transcription factors are regulatory proteins that bind to DNA, either promoting or inhibiting the transcription of a gene.[71] Within the genome of Escherichia coli bacteria, for example, there exists a series of genes necessary for the synthesis of the amino acid tryptophan. However, when tryptophan is already available to the jail cell, these genes for tryptophan synthesis are no longer needed. The presence of tryptophan directly affects the activeness of the genes—tryptophan molecules bind to the tryptophan repressor (a transcription factor), changing the repressor's structure such that the repressor binds to the genes. The tryptophan repressor blocks the transcription and expression of the genes, thereby creating negative feedback regulation of the tryptophan synthesis process.[72]

Transcription factors bind to Dna, influencing the transcription of associated genes.
Differences in cistron expression are especially clear inside multicellular organisms, where cells all contain the same genome but have very different structures and behaviors due to the expression of unlike sets of genes. All the cells in a multicellular organism derive from a single cell, differentiating into variant cell types in response to external and intercellular signals and gradually establishing different patterns of cistron expression to create dissimilar behaviors. As no single cistron is responsible for the development of structures inside multicellular organisms, these patterns ascend from the complex interactions betwixt many cells.
Within eukaryotes, at that place exist structural features of chromatin that influence the transcription of genes, often in the form of modifications to DNA and chromatin that are stably inherited by girl cells.[73] These features are chosen "epigenetic" considering they exist "on top" of the DNA sequence and retain inheritance from ane cell generation to the adjacent. Considering of epigenetic features, different cell types grown within the same medium can retain very different properties. Although epigenetic features are generally dynamic over the course of development, some, like the phenomenon of paramutation, have multigenerational inheritance and exist every bit rare exceptions to the general rule of Deoxyribonucleic acid every bit the basis for inheritance.[74]
Genetic change [edit]
Mutations [edit]

Gene duplication allows diversification past providing redundancy: one gene can mutate and lose its original office without harming the organism.
During the procedure of DNA replication, errors occasionally occur in the polymerization of the 2nd strand. These errors, called mutations, can affect the phenotype of an organism, especially if they occur within the protein coding sequence of a gene. Mistake rates are commonly very low—ane mistake in every 10–100 million bases—due to the "proofreading" ability of DNA polymerases.[75] [76] Processes that increase the rate of changes in Deoxyribonucleic acid are called mutagenic: mutagenic chemicals promote errors in DNA replication, often by interfering with the structure of base-pairing, while UV radiation induces mutations by causing harm to the DNA construction.[77] Chemical damage to Dna occurs naturally likewise and cells utilise DNA repair mechanisms to repair mismatches and breaks. The repair does not, however, e'er restore the original sequence. A especially important source of Deoxyribonucleic acid damages appears to be reactive oxygen species[78] produced by cellular aerobic respiration, and these can atomic number 82 to mutations.[79]
In organisms that use chromosomal crossover to exchange Deoxyribonucleic acid and recombine genes, errors in alignment during meiosis tin also cause mutations.[80] Errors in crossover are particularly probable when similar sequences cause partner chromosomes to prefer a mistaken alignment; this makes some regions in genomes more prone to mutating in this way. These errors create large structural changes in Dna sequence—duplications, inversions, deletions of unabridged regions—or the adventitious exchange of whole parts of sequences betwixt dissimilar chromosomes (chromosomal translocation).

This is a diagram showing mutations in an RNA sequence. Figure (1) is a normal RNA sequence, consisting of 4 codons. Figure (2) shows a missense, single indicate, not silent mutation. Figures (iii and 4) both show frameshift mutations, which is why they are grouped together. Figure iii shows a deletion of the 2nd base pair in the 2nd codon. Figure iv shows an insertion in the tertiary base of operations pair of the 2nd codon. Figure (five) shows a repeat expansion, where an entire codon is duplicated.
Natural choice and development [edit]
Mutations alter an organism's genotype and occasionally this causes different phenotypes to appear. Most mutations have niggling consequence on an organism's phenotype, wellness, or reproductive fettle.[81] Mutations that exercise take an issue are unremarkably detrimental, simply occasionally some can be benign.[82] Studies in the fly Drosophila melanogaster propose that if a mutation changes a protein produced by a gene, about 70 percent of these mutations volition be harmful with the remainder beingness either neutral or weakly benign.[83]

Population genetics studies the distribution of genetic differences inside populations and how these distributions change over time.[84] Changes in the frequency of an allele in a population are mainly influenced by natural selection, where a given allele provides a selective or reproductive advantage to the organism,[85] as well as other factors such as mutation, genetic drift, genetic hitchhiking,[86] artificial pick and migration.[87]
Over many generations, the genomes of organisms can modify significantly, resulting in development. In the procedure called accommodation, choice for beneficial mutations tin can crusade a species to evolve into forms better able to survive in their environment.[88] New species are formed through the process of speciation, often caused past geographical separations that prevent populations from exchanging genes with each other.[89]
By comparing the homology betwixt different species' genomes, it is possible to summate the evolutionary distance between them and when they may have diverged. Genetic comparisons are generally considered a more accurate method of characterizing the relatedness between species than the comparison of phenotypic characteristics. The evolutionary distances between species tin can be used to class evolutionary trees; these trees represent the mutual descent and divergence of species over fourth dimension, although they practice not evidence the transfer of genetic fabric between unrelated species (known equally horizontal gene transfer and most common in bacteria).[ninety]
Model organisms [edit]

Although geneticists originally studied inheritance in a wide range of organisms, researchers began to specialize in studying the genetics of a detail subset of organisms. The fact that pregnant enquiry already existed for a given organism would encourage new researchers to cull it for further study, and so somewhen a few model organisms became the footing for most genetics research.[91] Common research topics in model organism genetics include the study of cistron regulation and the involvement of genes in evolution and cancer.
Organisms were chosen, in part, for convenience—short generation times and piece of cake genetic manipulation made some organisms popular genetics research tools. Widely used model organisms include the gut bacterium Escherichia coli, the establish Arabidopsis thaliana, baker's yeast (Saccharomyces cerevisiae), the nematode Caenorhabditis elegans, the common fruit wing (Drosophila melanogaster), and the common house mouse (Mus musculus).
Medicine [edit]
Medical genetics seeks to sympathize how genetic variation relates to human health and disease.[92] When searching for an unknown gene that may be involved in a disease, researchers commonly use genetic linkage and genetic pedigree charts to find the location on the genome associated with the illness. At the population level, researchers take advantage of Mendelian randomization to look for locations in the genome that are associated with diseases, a method peculiarly useful for multigenic traits not clearly defined by a single gene.[93] Once a candidate gene is found, further enquiry is ofttimes done on the respective (or homologous) genes of model organisms. In improver to studying genetic diseases, the increased availability of genotyping methods has led to the field of pharmacogenetics: the study of how genotype can affect drug responses.[94]
Individuals differ in their inherited trend to develop cancer,[95] and cancer is a genetic disease.[96] The process of cancer development in the torso is a combination of events. Mutations occasionally occur within cells in the body as they divide. Although these mutations will not exist inherited past whatsoever offspring, they can bear on the beliefs of cells, sometimes causing them to grow and divide more frequently. There are biological mechanisms that attempt to finish this process; signals are given to inappropriately dividing cells that should trigger prison cell expiry, but sometimes additional mutations occur that cause cells to ignore these messages. An internal process of natural selection occurs within the body and eventually mutations accumulate within cells to promote their own growth, creating a cancerous tumor that grows and invades various tissues of the body.
Normally, a cell divides just in response to signals called growth factors and stops growing once in contact with surrounding cells and in response to growth-inhibitory signals. It unremarkably and so divides a limited number of times and dies, staying within the epithelium where it is unable to migrate to other organs. To become a cancer cell, a cell has to accumulate mutations in a number of genes (three to seven). A cancer cell can split without growth gene and ignores inhibitory signals. Likewise, it is immortal and tin abound indefinitely, even afterwards it makes contact with neighboring cells. It may escape from the epithelium and ultimately from the primary tumor. So, the escaped cell tin cantankerous the endothelium of a blood vessel and get transported by the bloodstream to colonize a new organ, forming deadly metastasis. Although at that place are some genetic predispositions in a small fraction of cancers, the major fraction is due to a set up of new genetic mutations that originally appear and accumulate in one or a small number of cells that will carve up to form the tumor and are non transmitted to the progeny (somatic mutations). The virtually frequent mutations are a loss of function of p53 protein, a tumor suppressor, or in the p53 pathway, and proceeds of function mutations in the Ras proteins, or in other oncogenes.
Research methods [edit]

DNA can be manipulated in the laboratory. Brake enzymes are commonly used enzymes that cutting Dna at specific sequences, producing anticipated fragments of DNA.[97] Deoxyribonucleic acid fragments tin be visualized through use of gel electrophoresis, which separates fragments according to their length.
The utilise of ligation enzymes allows Deoxyribonucleic acid fragments to be continued. By binding ("ligating") fragments of DNA together from unlike sources, researchers can create recombinant DNA, the DNA ofttimes associated with genetically modified organisms. Recombinant DNA is commonly used in the context of plasmids: brusque circular DNA molecules with a few genes on them. In the procedure known every bit molecular cloning, researchers can dilate the Dna fragments past inserting plasmids into leaner and then culturing them on plates of agar (to isolate clones of bacteria cells—"cloning" tin can as well refer to the various ways of creating cloned ("clonal") organisms).
DNA can as well be amplified using a procedure called the polymerase chain reaction (PCR).[98] By using specific brusk sequences of Dna, PCR tin can isolate and exponentially amplify a targeted region of Deoxyribonucleic acid. Because information technology can dilate from extremely small-scale amounts of Dna, PCR is besides oft used to detect the presence of specific Dna sequences.
Deoxyribonucleic acid sequencing and genomics [edit]
DNA sequencing, one of the almost fundamental technologies developed to study genetics, allows researchers to determine the sequence of nucleotides in DNA fragments. The technique of chain-termination sequencing, developed in 1977 by a team led by Frederick Sanger, is however routinely used to sequence DNA fragments.[99] Using this engineering, researchers have been able to report the molecular sequences associated with many homo diseases.
As sequencing has go less expensive, researchers have sequenced the genomes of many organisms using a process called genome assembly, which utilizes computational tools to run up together sequences from many different fragments.[100] These technologies were used to sequence the human genome in the Human Genome Project completed in 2003.[35] New high-throughput sequencing technologies are dramatically lowering the cost of Dna sequencing, with many researchers hoping to bring the cost of resequencing a man genome down to a grand dollars.[101]
Side by side-generation sequencing (or high-throughput sequencing) came about due to the e'er-increasing need for low-price sequencing. These sequencing technologies allow the production of potentially millions of sequences concurrently.[102] [103] The big amount of sequence information available has created the subfield of genomics, inquiry that uses computational tools to search for and analyze patterns in the total genomes of organisms. Genomics can too be considered a subfield of bioinformatics, which uses computational approaches to clarify large sets of biological data. A common problem to these fields of enquiry is how to manage and share data that deals with man subject and personally identifiable information.
Society and civilization [edit]
On nineteen March 2015, a group of leading biologists urged a worldwide ban on clinical use of methods, particularly the use of CRISPR and zinc finger, to edit the man genome in a way that tin can be inherited.[104] [105] [106] [107] In April 2015, Chinese researchers reported results of bones research to edit the DNA of non-viable human embryos using CRISPR.[108] [109]
Meet besides [edit]
- Bacterial genome size
- Cryoconservation of animal genetic resources
- Eugenics
- Embryology
- Development
- Genetic disorder
- Genetic multifariousness
- Genetic engineering
- Genetic enhancement
- Index of genetics articles
- Medical genetics
- Molecular tools for factor study
- Mutation
- Neuroepigenetics
- Outline of genetics
- Timeline of the history of genetics
- Plant genetic resources
References [edit]
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Genetics and the Organism: Introduction". An Introduction to Genetic Assay (seventh ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Hartl D, Jones Due east (2005)
- ^ "the definition of genetics". world wide web.dictionary.com . Retrieved 25 Oct 2018.
- ^ "Genetikos (γενετ-ικός)". Henry George Liddell, Robert Scott, A Greek-English language Lexicon. Perseus Digital Library, Tufts University. Archived from the original on 15 June 2010. Retrieved 20 February 2012.
- ^ "Genesis (γένεσις)". Henry George Liddell, Robert Scott, A Greek-English Dictionary. Perseus Digital Library, Tufts University. Archived from the original on 15 June 2010. Retrieved 20 Feb 2012.
- ^ "Genetic". Online Etymology Dictionary. Archived from the original on 23 Baronial 2011. Retrieved 20 February 2012.
- ^ Scientific discipline: The Definitive Visual Guide. Penguin. 2009. p. 362. ISBN978-0-7566-6490-9.
- ^ Weiling F (July 1991). "Historical report: Johann Gregor Mendel 1822–1884". American Periodical of Medical Genetics. forty (1): one–25, discussion 26. doi:10.1002/ajmg.1320400103. PMID 1887835.
- ^ Poczai P, Bong North, Hyvönen J (January 2014). "Imre Festetics and the Sheep Breeders' Society of Moravia: Mendel's Forgotten "Enquiry Network"". PLOS Biology. 12 (one): e1001772. doi:10.1371/journal.pbio.1001772. PMC3897355. PMID 24465180.
- ^ Hamilton Grand (2011). Population Genetics. Georgetown Academy. p. 26. ISBN978-1-4443-6245-9.
- ^ Lamarck, J-B (2008). In Encyclopædia Britannica. Retrieved from Encyclopædia Britannica Online on xvi March 2008.
- ^ Peter J. Bowler, The Mendelian Revolution: The Emergency of Hereditarian Concepts in Modern Science and Social club (Baltimore: Johns Hopkins University Press, 1989): chapters 2 & 3.
- ^ a b Blumberg RB. "Mendel's Paper in English". Archived from the original on thirteen January 2016.
- ^ genetics, n., Oxford English Lexicon, 3rd ed.
- ^ Bateson Due west. "Letter from William Bateson to Alan Sedgwick in 1905". The John Innes Centre. Archived from the original on 13 October 2007. Retrieved 15 March 2008. Note that the letter was to an Adam Sedgwick, a zoologist and "Reader in Animal Morphology" at Trinity College, Cambridge
- ^ genetic, adj., Oxford English Dictionary, 3rd ed.
- ^ Richmond ML (November 2007). "Opportunities for women in early genetics". Nature Reviews Genetics. viii (eleven): 897–902. doi:x.1038/nrg2200. PMID 17893692. S2CID 21992183. Archived from the original on 16 May 2008.
- ^ Bateson W (1907). "The Progress of Genetic Research". In Wilks, West (ed.). Report of the Third 1906 International Conference on Genetics: Hybridization (the cross-breeding of genera or species), the cross-breeding of varieties, and full general plant breeding. London: Royal Horticultural Order. :Initially titled the "International Conference on Hybridisation and Plant Convenance", the title was changed as a result of Bateson's speech. See: Cock AG, Forsdyke DR (2008). Treasure your exceptions: the science and life of William Bateson . Springer. p. 248. ISBN978-0-387-75687-5.
- ^ a b c "Nettie Stevens: A Discoverer of Sex Chromosomes". Scitable. Nature Education. Retrieved 8 June 2020.
- ^ Moore, John A. (1983). "Thomas Hunt Morgan – The Geneticist". Integrative and Comparative Biological science. 23 (iv): 855–65. doi:10.1093/icb/23.4.855.
- ^ Sturtevant AH (1913). "The linear arrangement of half-dozen sexual practice-linked factors in Drosophila, equally shown by their way of association" (PDF). Journal of Experimental Biology. xiv: 43–59. CiteSeerXx.1.1.37.9595. doi:x.1002/jez.1400140104. Archived (PDF) from the original on 27 February 2008.
- ^ Avery OT, Macleod CM, McCarty M (February 1944). "Studies on the Chemical Nature of the Substance Inducing Transformation of Pneumococcal Types: Induction of Transformation past a Desoxyribonucleic Acid Fraction Isolated from Pneumococcus Type Three". The Journal of Experimental Medicine. 79 (2): 137–58. doi:10.1084/jem.79.ii.137. PMC2135445. PMID 19871359. Reprint: Avery OT, MacLeod CM, McCarty 1000 (February 1979). "Studies on the chemical nature of the substance inducing transformation of pneumococcal types. Inductions of transformation by a desoxyribonucleic acid fraction isolated from pneumococcus blazon Iii". The Periodical of Experimental Medicine. 149 (two): 297–326. doi:10.1084/jem.149.2.297. PMC2184805. PMID 33226.
- ^ Khanna P (2008). Prison cell and Molecular Biology. I.K. International Pvt Ltd. p. 221. ISBN978-81-89866-59-four.
- ^ Hershey Advert, Chase 1000 (May 1952). "Independent functions of viral protein and nucleic acid in growth of bacteriophage". The Periodical of General Physiology. 36 (1): 39–56. doi:10.1085/jgp.36.1.39. PMC2147348. PMID 12981234.
- ^ Judson H (1979). The Eighth Day of Creation: Makers of the Revolution in Biological science. Cold Spring Harbor Laboratory Press. pp. 51–169. ISBN978-0-87969-477-7.
- ^ Watson JD, Crick FH (Apr 1953). "Molecular construction of nucleic acids; a construction for deoxyribose nucleic acid" (PDF). Nature. 171 (4356): 737–8. Bibcode:1953Natur.171..737W. doi:10.1038/171737a0. PMID 13054692. S2CID 4253007. Archived (PDF) from the original on 4 Feb 2007.
- ^ Watson JD, Crick FH (May 1953). "Genetical implications of the structure of deoxyribonucleic acid" (PDF). Nature. 171 (4361): 964–7. Bibcode:1953Natur.171..964W. doi:10.1038/171964b0. PMID 13063483. S2CID 4256010. Archived (PDF) from the original on 21 June 2003.
- ^ Stratmann SA, van Oijen AM (February 2014). "Deoxyribonucleic acid replication at the single-molecule level" (PDF). Chemical Society Reviews. 43 (4): 1201–20. doi:x.1039/c3cs60391a. PMID 24395040. S2CID 205856075.
- ^ Betz F (2010). Managing Scientific discipline: Methodology and Organization of Research. Springer. p. 76. ISBN978-1-4419-7488-four.
- ^ Rice SA (2009). Encyclopedia of Development. Infobase Publishing. p. 134. ISBN978-1-4381-1005-9.
- ^ Sarkar S (1998). Genetics and Reductionism. Cambridge Academy Press. p. 140. ISBN978-0-521-63713-8.
- ^ Ohta T (November 1973). "Slightly deleterious mutant substitutions in development". Nature. 246 (5428): 96–eight. Bibcode:1973Natur.246...96O. doi:10.1038/246096a0. PMID 4585855. S2CID 4226804.
- ^ Sanger F, Nicklen South, Coulson AR (December 1977). "Dna sequencing with chain-terminating inhibitors". Proceedings of the National Academy of Sciences of the Usa of America. 74 (12): 5463–7. Bibcode:1977PNAS...74.5463S. doi:10.1073/pnas.74.12.5463. PMC431765. PMID 271968.
- ^ Saiki RK, Scharf Due south, Faloona F, Mullis KB, Horn GT, Erlich HA, Arnheim North (Dec 1985). "Enzymatic distension of beta-globin genomic sequences and brake site analysis for diagnosis of sickle cell anemia". Science. 230 (4732): 1350–iv. Bibcode:1985Sci...230.1350S. doi:x.1126/scientific discipline.2999980. PMID 2999980.
- ^ a b "Human Genome Project Information". Human Genome Project. Archived from the original on xv March 2008. Retrieved fifteen March 2008.
- ^ "The sequence of the human genome". Science. 291.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Patterns of Inheritance: Introduction". An Introduction to Genetic Analysis (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Mendel'southward experiments". An Introduction to Genetic Analysis (7th ed.). New York: Westward.H. Freeman. ISBN978-0-7167-3520-v.
- ^ a b c Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Mendelian genetics in eukaryotic life cycles". An Introduction to Genetic Assay (7th ed.). New York: West.H. Freeman. ISBN978-0-7167-3520-five.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Interactions between the alleles of ane gene". An Introduction to Genetic Analysis (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Cheney RW. "Genetic Notation". Christopher Newport University. Archived from the original on three January 2008. Retrieved eighteen March 2008.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Human Genetics". An Introduction to Genetic Analysis (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Cistron interaction and modified dihybrid ratios". An Introduction to Genetic Assay (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-v.
- ^ Mayeux R (June 2005). "Mapping the new frontier: complex genetic disorders". The Journal of Clinical Investigation. 115 (half dozen): 1404–seven. doi:x.1172/JCI25421. PMC1137013. PMID 15931374.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Quantifying heritability". An Introduction to Genetic Assay (7th ed.). New York: Due west. H. Freeman. ISBN978-0-7167-3520-v.
- ^ Luke A, Guo Ten, Adeyemo AA, Wilks R, Forrester T, Lowe W, et al. (July 2001). "Heritability of obesity-related traits among Nigerians, Jamaicans and US black people". International Journal of Obesity and Related Metabolic Disorders. 25 (seven): 1034–41. doi:10.1038/sj.ijo.0801650. PMID 11443503.
- ^ Pearson H (May 2006). "Genetics: what is a cistron?". Nature. 441 (7092): 398–401. Bibcode:2006Natur.441..398P. doi:ten.1038/441398a. PMID 16724031. S2CID 4420674.
- ^ Prescott, L (1993). Microbiology. Wm. C. Brown Publishers. ISBN978-0-697-01372-9.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Machinery of Dna Replication". An Introduction to Genetic Analysis (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-v.
- ^ Gregory SG, Barlow KF, McLay KE, Kaul R, Swarbreck D, Dunham A, et al. (May 2006). "The DNA sequence and biological annotation of human chromosome i". Nature. 441 (7091): 315–21. Bibcode:2006Natur.441..315G. doi:10.1038/nature04727. PMID 16710414.
- ^ Alberts et al. (2002), II.4. Deoxyribonucleic acid and chromosomes: Chromosomal DNA and Its Packaging in the Chromatin Fiber Archived 18 October 2007 at the Wayback Machine
- ^ a b c "Ruth Sager". Encyclopaedia Britannica . Retrieved 8 June 2020.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Sex chromosomes and sexual practice-linked inheritance". An Introduction to Genetic Analysis (seventh ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-five.
- ^ a b Rastan, Sohaila (2015). "Mary F. Lyon (1925–2014)". Nature. Springer Nature Limited. 518 (7537): 36. Bibcode:2015Natur.518...36R. doi:x.1038/518036a. PMID 25652989. S2CID 4405984.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Bacterial conjugation". An Introduction to Genetic Analysis (7th ed.). New York: Due west.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Bacterial transformation". An Introduction to Genetic Analysis (seventh ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-v.
- ^ a b c Bernstein H, Bernstein C, Michod RE (2018). "Sex in microbial pathogens". Infect Genet Evol. 57: 8–25. doi:10.1016/j.meegid.2017.10.024. PMID 29111273.
{{cite periodical}}: CS1 maint: multiple names: authors list (link) - ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Nature of crossing-over". An Introduction to Genetic Analysis (7th ed.). New York: W. H. Freeman. ISBN978-0-7167-3520-v.
- ^ Creighton H. B. (1931). "A Correlation of Cytological and Genetical Crossing-Over in Zea Mays". Proc Natl Acad Sci United states. 17 (8): 492–497. Bibcode:1931PNAS...17..492C. doi:10.1073/pnas.17.8.492. PMC1076098. PMID 16587654.
- ^ Staub JE (1994). Crossover: Concepts and Applications in Genetics, Evolution, and Breeding. Academy of Wisconsin Press. p. 55. ISBN978-0-299-13564-5.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Linkage maps". An Introduction to Genetic Analysis (seventh ed.). New York: West. H. Freeman. ISBN978-0-7167-3520-5.
- ^ Berg JM, Tymoczko JL, Stryer Fifty, Clarke ND (2002). "I. 5. Deoxyribonucleic acid, RNA, and the Menses of Genetic Information: Amino Acids Are Encoded past Groups of Three Bases Starting from a Fixed Indicate". Biochemistry (5th ed.). New York: W.H. Freeman and Company. Archived from the original on 11 Apr 2006.
- ^ Crick F (Baronial 1970). "Cardinal dogma of molecular biology" (PDF). Nature. 227 (5258): 561–3. Bibcode:1970Natur.227..561C. doi:ten.1038/227561a0. PMID 4913914. S2CID 4164029. Archived (PDF) from the original on 15 Feb 2006.
- ^ Alberts et al. (2002), I.3. Proteins: The Shape and Structure of Proteins
- ^ Alberts et al. (2002), I.3. Proteins: Poly peptide Function Archived 25 April 2006 at the Wayback Motorcar
- ^ "How Does Sickle Prison cell Cause Disease?". Brigham and Women'southward Infirmary: Information Middle for Sickle Cell and Thalassemic Disorders. 11 April 2002. Archived from the original on 23 September 2010. Retrieved 23 July 2007.
- ^ Imes DL, Geary LA, Grahn RA, Lyons LA (April 2006). "Albinism in the domestic true cat (Felis catus) is associated with a tyrosinase (TYR) mutation". Animal Genetics. 37 (2): 175–8. doi:ten.1111/j.1365-2052.2005.01409.10. PMC1464423. PMID 16573534.
- ^ "MedlinePlus: Phenylketonuria". NIH: National Library of Medicine. Archived from the original on 25 July 2008. Retrieved 15 March 2008.
- ^ For instance, Ridley M (2003). Nature via Nurture: Genes, Experience and What Makes Us Human. Fourth Manor. p. 73. ISBN978-1-84115-745-0.
- ^ Rosenthal D (1964). "The Genain Quadruplets: A Case Study and Theoretical Analysis of Heredity and Surroundings in Schizophrenia". Behavioral Science. 9 (four): 371. doi:x.1002/bs.3830090407.
- ^ Brivanlou AH, Darnell JE (February 2002). "Signal transduction and the control of gene expression". Scientific discipline. 295 (5556): 813–viii. Bibcode:2002Sci...295..813B. CiteSeerXten.1.1.485.6042. doi:x.1126/science.1066355. PMID 11823631. S2CID 14954195.
- ^ Alberts et al. (2002), 2.three. Control of Gene Expression – The Tryptophan Repressor is a Simple Switch That Turns Genes On and Off in Bacteria Archived 29 June 2007 at the Wayback Machine
- ^ Jaenisch R, Bird A (March 2003). "Epigenetic regulation of factor expression: how the genome integrates intrinsic and environmental signals". Nature Genetics. 33: 245–54. doi:10.1038/ng1089. PMID 12610534. S2CID 17270515.
- ^ Chandler VL (February 2007). "Paramutation: from maize to mice". Cell. 128 (4): 641–five. doi:10.1016/j.cell.2007.02.007. PMID 17320501. S2CID 6928707.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Spontaneous mutations". An Introduction to Genetic Analysis (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Freisinger E, Grollman AP, Miller H, Kisker C (April 2004). "Lesion (in)tolerance reveals insights into DNA replication fidelity". The EMBO Periodical. 23 (7): 1494–505. doi:10.1038/sj.emboj.7600158. PMC391067. PMID 15057282.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Induced mutations". An Introduction to Genetic Analysis (7th ed.). New York: W. H. Freeman. ISBN978-0-7167-3520-5.
- ^ Cadet J, Wagner JR (February 2013). "Dna base of operations harm by reactive oxygen species, oxidizing agents, and UV radiation". Cold Spring Harbor Perspectives in Biology. 5 (2): a012559. doi:10.1101/cshperspect.a012559. PMC3552502. PMID 23378590.
- ^ Jena NR (July 2012). "DNA damage past reactive species: Mechanisms, mutation and repair". Journal of Biosciences. 37 (3): 503–17. doi:10.1007/s12038-012-9218-2. PMID 22750987. S2CID 14837181.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Chromosome Mutation I: Changes in Chromosome Structure: Introduction". An Introduction to Genetic Assay (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Schaechter M (2009). Encyclopedia of Microbiology. Bookish Printing. p. 551. ISBN978-0-12-373944-5.
- ^ Calver Grand, Lymbery A, McComb J, Bamford M (2009). Ecology Biology. Cambridge University Press. p. 118. ISBN978-0-521-67982-iv.
- ^ Sawyer SA, Parsch J, Zhang Z, Hartl DL (April 2007). "Prevalence of positive selection among about neutral amino acid replacements in Drosophila". Proceedings of the National Academy of Sciences of the United states of america. 104 (xvi): 6504–10. Bibcode:2007PNAS..104.6504S. doi:ten.1073/pnas.0701572104. PMC1871816. PMID 17409186.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Variation and its modulation". An Introduction to Genetic Analysis (7th ed.). New York: Westward.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Pick". An Introduction to Genetic Analysis (7th ed.). New York: W. H. Freeman. ISBN978-0-7167-3520-5.
- ^ Gillespie JH (November 2001). "Is the population size of a species relevant to its development?". Development; International Journal of Organic Development. 55 (xi): 2161–9. doi:10.1111/j.0014-3820.2001.tb00732.10. PMID 11794777. S2CID 221735887.
- ^ Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). "Random events". An Introduction to Genetic Assay (7th ed.). New York: W.H. Freeman. ISBN978-0-7167-3520-5.
- ^ Darwin C (1859). On the Origin of Species (1st ed.). London: John Murray. p. 1. ISBN978-0-8014-1319-3. Archived from the original on 12 December 2006.
Earlier related ideas were best-selling in Darwin C (1861). On the Origin of Species (3rd ed.). London: John Murray. xiii. ISBN978-0-8014-1319-3. Archived from the original on 23 February 2011. - ^ Gavrilets S (October 2003). "Perspective: models of speciation: what have we learned in twoscore years?". Development; International Periodical of Organic Evolution. 57 (10): 2197–215. doi:10.1554/02-727. PMID 14628909. S2CID 198158082.
- ^ Wolf YI, Rogozin IB, Grishin NV, Koonin EV (September 2002). "Genome trees and the tree of life". Trends in Genetics. eighteen (9): 472–ix. doi:10.1016/S0168-9525(02)02744-0. PMID 12175808.
- ^ "The Use of Model Organisms in Instruction". University of Wisconsin: Wisconsin Outreach Research Modules. Archived from the original on thirteen March 2008. Retrieved xv March 2008.
- ^ "NCBI: Genes and Illness". NIH: National Heart for Biotechnology Data. Archived from the original on xx February 2007. Retrieved fifteen March 2008.
- ^ Smith GD, Ebrahim S (February 2003). "'Mendelian randomization': can genetic epidemiology contribute to agreement environmental determinants of disease?". International Journal of Epidemiology. 32 (1): ane–22. doi:ten.1093/ije/dyg070. PMID 12689998.
- ^ "Pharmacogenetics Fact Sail". NIH: National Institute of General Medical Sciences. Archived from the original on 12 May 2008. Retrieved 15 March 2008.
- ^ Frank SA (October 2004). "Genetic predisposition to cancer – insights from population genetics". Nature Reviews Genetics. v (10): 764–72. doi:x.1038/nrg1450. PMID 15510167. S2CID 6049662.
- ^ Strachan T, Read AP (1999). Human Molecular Genetics two (second ed.). John Wiley & Sons Inc. Chapter 18: Cancer Genetics Archived 26 September 2005 at the Wayback Car
- ^ Lodish et al. (2000), Affiliate 7: 7.ane. Deoxyribonucleic acid Cloning with Plasmid Vectors Archived 27 May 2009 at the Wayback Machine
- ^ Lodish et al. (2000), Affiliate 7: 7.seven. Polymerase Chain Reaction: An Alternative to Cloning
- ^ Brown TA (2002). "Section ii, Chapter 6: 6.i. The Methodology for DNA Sequencing". Genomes 2 (2nd ed.). Oxford: Bios. ISBN978-1-85996-228-2.
- ^ Brown (2002), Section 2, Chapter half-dozen: half dozen.2. Associates of a Face-to-face DNA Sequence Archived eight Feb 2007 at the Wayback Motorcar
- ^ Service RF (March 2006). "Gene sequencing. The race for the $1000 genome". Science. 311 (5767): 1544–half-dozen. doi:10.1126/science.311.5767.1544. PMID 16543431. S2CID 23411598.
- ^ Hall N (May 2007). "Avant-garde sequencing technologies and their wider touch on in microbiology". The Journal of Experimental Biology. 210 (Pt 9): 1518–25. doi:10.1242/jeb.001370. PMID 17449817.
- ^ Church GM (January 2006). "Genomes for all". Scientific American. 294 (1): 46–54. Bibcode:2006SciAm.294a..46C. doi:ten.1038/scientificamerican0106-46. PMID 16468433. (subscription required)
- ^ Wade N (nineteen March 2015). "Scientists Seek Ban on Method of Editing the Man Genome". The New York Times. Archived from the original on 19 March 2015. Retrieved 20 March 2015.
- ^ Pollack A (3 March 2015). "A Powerful New Way to Edit Deoxyribonucleic acid". The New York Times. Archived from the original on 26 March 2015. Retrieved 20 March 2015.
- ^ Baltimore D, Berg P, Botchan 1000, Carroll D, Charo RA, Church building G, et al. (April 2015). "Biotechnology. A prudent path forrad for genomic engineering science and germline factor modification". Science. 348 (6230): 36–viii. Bibcode:2015Sci...348...36B. doi:10.1126/science.aab1028. PMC4394183. PMID 25791083.
- ^ Lanphier E, Urnov F, Haecker SE, Werner One thousand, Smolenski J (March 2015). "Don't edit the homo germ line". Nature. 519 (7544): 410–ane. Bibcode:2015Natur.519..410L. doi:10.1038/519410a. PMID 25810189.
- ^ Kolata M (23 April 2015). "Chinese Scientists Edit Genes of Human being Embryos, Raising Concerns". The New York Times. Archived from the original on 24 April 2015. Retrieved 24 April 2015.
- ^ Liang P, Xu Y, Zhang X, Ding C, Huang R, Zhang Z, et al. (May 2015). "CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes". Protein & Jail cell. 6 (5): 363–372. doi:10.1007/s13238-015-0153-five. PMC4417674. PMID 25894090.
Farther reading [edit]
- Alberts B, Bray D, Hopkin M, Johnson A, Lewis J, Raff M, Roberts M, Walter P (2013). Essential Cell Biology, 4th Edition. Garland Science. ISBN978-1-317-80627-1.
- Griffiths AJ, Miller JH, Suzuki DT, Lewontin RC, Gelbart, eds. (2000). An Introduction to Genetic Analysis (seventh ed.). New York: W. H. Freeman. ISBN978-0-7167-3520-five.
- Hartl D, Jones E (2005). Genetics: Analysis of Genes and Genomes (sixth ed.). Jones & Bartlett. ISBN978-0-7637-1511-iii.
- Male monarch RC, Mulligan PK, Stansfield WD (2013). A Dictionary of Genetics (8th ed.). New York: Oxford University Press. ISBN978-0-19-976644-4.
- Lodish H, Berk A, Zipursky LS, Matsudaira P, Baltimore D, Darnell J (2000). Molecular Prison cell Biological science (4th ed.). New York: Scientific American Books. ISBN978-0-7167-3136-8.
External links [edit]
| | Wikiquote has quotations related to: Genetics |
- Genetics on In Our Time at the BBC
- Genetics at Curlie
Source: https://en.wikipedia.org/wiki/Genetics
Mag-post ng isang Komento for "What is the term used to describe the scientific study of heredity?"